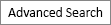- Search
| Ann Coloproctol > Volume 40(1); 2024 > Article |
|
Abstract
Purpose
Methods
Results
Conclusion
Notes
Conflict of interest
Bo Young Oh and Il Tae Son are Editorial Board members of Annals of Coloproctology, but were not involved in in the peer reviewer selection, evaluation, or decision process of this article. No other potential conflict of interest relevant to this article was reported.
Author contributions
Conceptualization: ITS, BJC; Formal analysis: MK, PT; Investigation: MK, PT, MJK, BYO; Methodology: PT; Project administration: ITS, BJC; Supervision: MJK, BYO, Validation: MJK, BYO, WritingŌĆōoriginal draft: MK; WritingŌĆōreview & editing: all authors. All authors read and approved the final manuscript.
Fig.┬Ā1.

Fig.┬Ā2.
Table┬Ā1.
| Study | Country | Modality | Architecture | Internal dataset | External dataset | Automatization | Performance | Comparison |
|---|---|---|---|---|---|---|---|---|
| Tumor segmentation | ||||||||
| ŌĆāTrebeschi et al. [26] (2017) | The Netherlands | T2WI, DWI | CNN | Training, 56 | None | Manual annotation | DSC, 0.70 | None |
| Validation, 62 | AUROC, 0.99 | |||||||
| Test, 14 | ||||||||
| ŌĆāWang et al. [27] (2018) | China | T2WI | CNN (2D U-Net) | Training, 84 | None | Manual annotation | DSC, 0.74 | None |
| Validation, 9 | Hausdorff distance, 20.44 | |||||||
| Test, 20 | Average surface distance, 3.25 | |||||||
| Jaccard index, 0.60 | ||||||||
| ŌĆāKim et al. [28] (2019) | Korea | T2WI | CNN (U-Net, FCN-8, SegNet) | 133 | None | Manual annotation | U-Net: | U-Net is superior to other models |
| ŌĆāDSC, 0.81 | ||||||||
| ŌĆāSensitivity, 0.79 | ||||||||
| ŌĆāSpecificity, 0.98 | ||||||||
| ŌĆāPang et al. [29] (2021) | China | T2WI | CNN (U-Net) | Training, 88 | 34 | Manual annotation | DSC, 0.95 | None |
| Test, 46 | Sensitivity, 0.97 | |||||||
| Specificity, 0.96 | ||||||||
| ŌĆāKnuth et al. [30] (2022) | Norway | T2WI | CNN (2D U-Net) | 109 | 83 | Manual annotation | DSC, 0.78 | None |
| ŌĆāDeSilvio et al. [31] (2023) | USA | T2WI | CNN (region-specific U-Net, multiclass U-Net) | Training, 44 | 11 | Manual annotation | Region-specific U-Net: | Region-specific U-Net is superior to multiclass U-Net |
| Validation, 44 | ŌĆāDSC, 0.91 | |||||||
| Test, 49 | ŌĆāHausdorff distance, 2.45 | |||||||
| ŌĆāZhang et al. [32] (2021) | China | T2WI, DWI | CNN (3D V-Net) | Training, 108 | None | Manual annotation | DSC, 0.96 | None |
| Test, 94 | ||||||||
| ŌĆāJian et al. [34] (2018) | China | T2WI | CNN (VGG-16) | Training, 410 | None | Fully automatic | DSC, 0.84 | VGG-16 is superior to U-Net |
| Test, 102 | PPV, 0.83 | |||||||
| Sensitivity, 0.88 | ||||||||
| Specificity, 0.97 | ||||||||
| Hammoude distance, 0.27 | ||||||||
| Hausdorff distance, 8.2 | ||||||||
| ŌĆāZhu et al. [35] (2021) | China | DWI | CNN (3D U-Net) | Training, 180 | None | Fully automatic | DSC, 0.68 | Automatic model is superior to semiautomatic model |
| Validation, 60 | ||||||||
| Test, 60 | ||||||||
| LN segmentation | ||||||||
| ŌĆāZhao et al. [33] (2020) | China | T2WI, DWI | CNN (Mask R-CNN) | 293 | 50 | Manual annotation | Detection: | Mask R-CNN is superior to junior radiologists in LN detection |
| ŌĆāSensitivity, 0.63 | ||||||||
| ŌĆāPPV, 0.65 | ||||||||
| ŌĆāFalse-positive rate per case, 8.2 | ||||||||
| Segmentation: | ||||||||
| ŌĆāDSC, 0.81-0.82 |
T2WI, T2-weighted image; DWI, diffusion-weighted image; CNN, convolutional neural network; DSC, dice similarity coefficient; AUROC, area under the receiver operating characteristic curve; 2D, 2-dimensional; FCN-8, fully convolutional network (8 pixels); 3D, 3-dimensional; LN, lymph node; PPV, positive predictive value.
Table┬Ā2.
| Study | Country | Modality | Architecture | Internal dataset | External dataset | Performance | Comparison |
|---|---|---|---|---|---|---|---|
| T stage | |||||||
| ŌĆāKim et al. [28] (2019) | Korea | T2WI | CNN (AlexNet, Inception) | 133 | None | Inception: | Inception is superior to AlexNet |
| ŌĆāAccuracy, 0.94 | |||||||
| ŌĆāSensitivity, 0.88 | |||||||
| ŌĆāSpecificity, 1.0 | |||||||
| ŌĆāWu et al. [43] (2021) | China | T2WI | CNN (Faster R-CNN) | 183 | None | Coronal view: | None |
| ŌĆāAUROC (T1), 0.96 | |||||||
| ŌĆāAUROC (T2), 0.97 | |||||||
| ŌĆāAUROC (T3), 0.97 | |||||||
| ŌĆāAUROC (T4), 0.97 | |||||||
| N stage | |||||||
| ŌĆāLu et al. [44] (2018) | China | T2WI, DWI | CNN (Faster R-CNN) | 351 | 414 | AUROC, 0.91 | No difference in accuracy compared to radiologists |
| Diagnostic time, 20 sec vs. 600 sec | Faster than radiologists | ||||||
| ŌĆāDing et al. [45] (2019) | China | T2WI, DWI | CNN (Faster R-CNN) | 414 | None | ╬║ coefficient: | AI model is superior to radiologists |
| AI vs. pathologist, 0.57 | |||||||
| Radiologist vs. pathologist, 0.47 | |||||||
| ŌĆāZhou et al. [46] (2019) | China | Pelvic HD MRI | CNN | Training, 201 | None | AUROC, 0.89 | No difference in accuracy compared with radiologists |
| Test, 100 | Diagnostic time, 10 sec vs. 600 sec | Faster than radiologists | |||||
| ŌĆāLi et al. [47] (2021) | China | T2WI | CNN (InceptionV3) | 129 | None | Sensitivity, 0.95 | AI is superior to radiologists |
| Specificity, 0.95 | |||||||
| PPV, 0.95 | |||||||
| NPV, 0.95 | |||||||
| AUROC, 0.99 | |||||||
| Circumferential resection margin | |||||||
| ŌĆāWang et al. [48] (2020) | China | T2WI | CNN (Faster R-CNN) | Training, 192 | None | Accuracy, 0.93 | None |
| Test, 48 | Sensitivity, 0.84 | ||||||
| Specificity, 0.96 | |||||||
| AUROC, 0.95 | |||||||
| ŌĆāXu et al. [49] (2020) | China | T2WI | CNN (Faster R-CNN) | Training, 300 | None | Accuracy, 0.88 | None |
| Test, 50 | Sensitivity, 0.86 | ||||||
| Specificity, 0.90 | |||||||
| AUROC, 0.93 | |||||||
Table┬Ā3.
| Study | Country | Modality | Architecture | Internal dataset | External dataset | Performance | Comparison |
|---|---|---|---|---|---|---|---|
| Microsatellite instability | |||||||
| ŌĆāZhang et al. [61] (2021) | China | T2WI | CNN (3D MobileNetV2) | Training, 395 | None | Image model: | Image model is superior to clinical model |
| Validation, 395 | ŌĆāSensitivity, 0.89 | ||||||
| Test, 96 | ŌĆāSpecificity, 0.74 | ||||||
| ŌĆāAUROC, 0.82 | |||||||
| Clinical model: | |||||||
| ŌĆāSensitivity, 1.00 | |||||||
| ŌĆāSpecificity, 0.31 | |||||||
| ŌĆāAUROC, 0.61 | |||||||
| ŌĆāCao et al. [62] (2023) | China | Enhanced APCT | CNN (Resnet101) | Training, 1,124 | 206 | Internal validation: | None |
| Test, 482 | ŌĆāAccuracy, 0.99 | ||||||
| ŌĆāSensitivity, 1.0 | |||||||
| ŌĆāSpecificity, 0.97 | |||||||
| ŌĆāAUROC, 0.99 | |||||||
| External validation: | |||||||
| ŌĆāAccuracy, 0.91 | |||||||
| ŌĆāSensitivity, 0.90 | |||||||
| ŌĆāSpecificity, 0.93 | |||||||
| ŌĆāAUROC, 0.92 | |||||||
| KRAS mutation | |||||||
| ŌĆāHe et al. [63] (2020) | China | Enhanced APCT | CNN (ResNet) | Training, 117 | None | AI model: | AI model is superior to radiomics model |
| Test, 40 | ŌĆāSensitivity, 0.59 | ||||||
| ŌĆāSpecificity, 1.0 | |||||||
| ŌĆāAUROC, 0.93 | |||||||
| Radiomics model: | |||||||
| ŌĆāSensitivity, 0.70 | |||||||
| ŌĆāSpecificity, 0.85 | |||||||
| ŌĆāAUROC, 0.82 | |||||||
Table┬Ā4.
| Study | Country | Modality | Architecture | Internal dataset | External dataset | Performance | Comparison |
|---|---|---|---|---|---|---|---|
| Shi et al. [79] (2019) | China | T2WI (pre- and mid-CRT) | CNN | 51 | None | pCR: | No difference in predicting pCR |
| ŌĆāAUROC (CNN), 0.83 | CNN model is inferior to radiomics model in predicting GR | ||||||
| ŌĆāAUROC (radiomics), 0.81 | |||||||
| GR: | |||||||
| ŌĆāAUROC (CNN), 0.74 | |||||||
| ŌĆāAUROC (radiomics), 0.92 | |||||||
| Zhang et al. [80] (2020) | China | T2WI (pre- and post-CRT) | CNN (models A, B, C) | Training, 290 | 93 | pCR: | Model A is superior to the other models |
| ŌĆāAccuracy, 0.98 | |||||||
| ŌĆāSensitivity, 1.00 | |||||||
| ŌĆāSpecificity, 0.97 | |||||||
| ŌĆāAUROC, 0.99 | |||||||
| Zhu et al. [81] (2020) | China | T2WI (pre-CRT) | CNN | Training, 400 | None | GR: | None |
| Validation, 100 | ŌĆāSensitivity, 0.93 | ||||||
| Test, 200 | ŌĆāSpecificity, 0.62 | ||||||
| ŌĆāAUROC, 0.81 | |||||||
| Jang et al. [78] (2021) | Korea | T2WI (post-CRT) | CNN (ShuffleNet) | Training, 303 | None | pCR: | AI model is superior to human for predicting pCR and GR |
| Validation, 46 | ŌĆāAccuracy, 0.85 | ||||||
| Test, 117 | ŌĆāSensitivity, 0.30 | ||||||
| ŌĆāSpecificity, 0.96 | |||||||
| GR: | |||||||
| ŌĆāAccuracy 0.72 | |||||||
| ŌĆāSensitivity 0.54 | |||||||
| ŌĆāSpecificity 0.81 |